Applying Tensor-based Morphometry to Parametric Surfaces can Improve MRI-based Disease Diagnosis
Yalin Wang, Lei Yuan, Jie Shi, Alexander Greve, Jieping Ye, Arthur W. Toga, Allan L. Reiss and Paul M. Thompson
Abstract
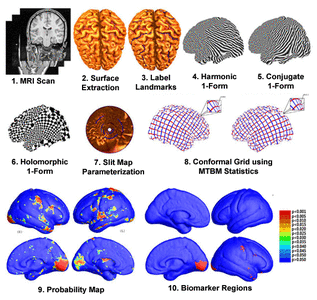
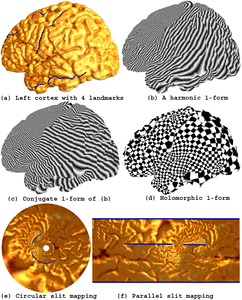
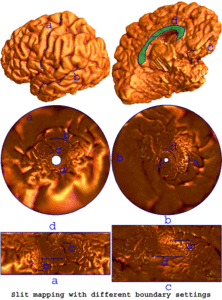
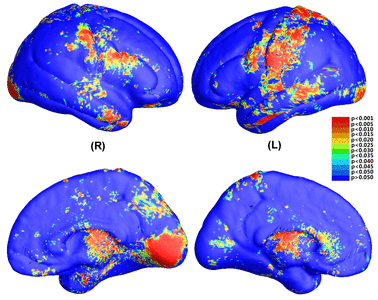
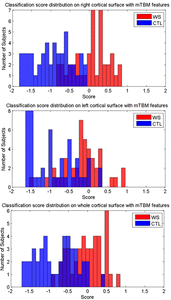
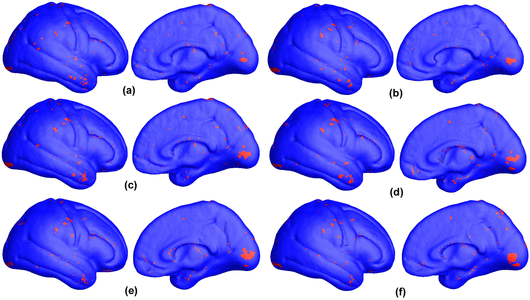
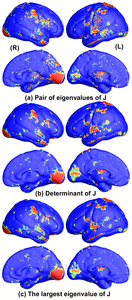
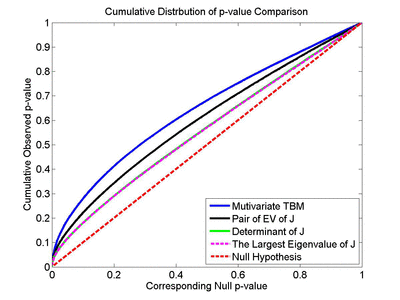
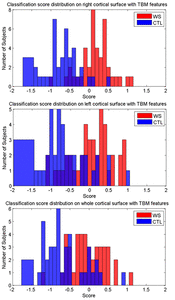
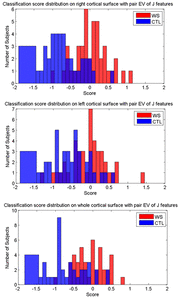
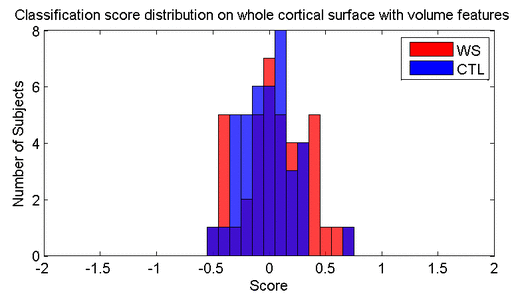
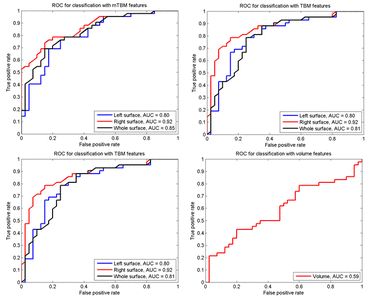
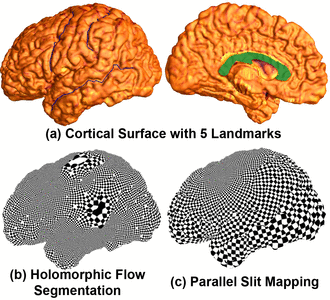
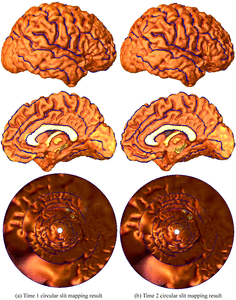
Many methods have been proposed for computer-assisted diagnostic classification. Full tensor information and machine learning with 3D maps derived from brain images may help detect subtle differences or classify subjects into different groups. Here we develop a new approach to apply tensor-based morphometry to parametric surface models for diagnostic classification. We use this approach to identify cortical surface features for use in diagnostic classifiers. First, with holomorphic 1-forms, we compute an efficient and accurate conformal mapping from a multiply connected mesh to the so-called slit domain. Next, the surface parameterization approach provides a natural way to register anatomical surfaces across subjects using a constrained harmonic map. To analyze anatomical differences, we then analyze the full Riemannian surface metric tensors, which retain multivariate information on local surface geometry. As the number of voxels in a 3D image is large, sparse learning is a promising method to select a subset of imaging features and to improve classification accuracy. Focusing on vertices with greatest effect sizes, we train a diagnostic classifier using the surface features selected by an L1-norm based sparse learning method. Stability selection is applied to validate the selected feature sets. We tested the algorithm on MRI-derived cortical surfaces from 42 subjects with genetically confirmed Williams syndrome and 40 age-matched controls, multivariate statistics on the local tensors gave greater effect sizes for detecting group differences relative to other TBM-based statistics including analysis of the Jacobian determinant and the largest eigenvalue of the surface metric. Our method also gave reasonable classification results relative to the Jacobian determinant, the pair of eigenvalues of the Jacobian matrix and volume features. This analysis pipeline may boost the power of morphometry studies, and may assist with image-based classification.
Figures (click on each for a larger version):
Related Publications
- Y. Wang, L. Yuan, J. Shi, A. Greve, J. Ye, A. W. Toga, A. L. Reiss and P. M. Thompson, Applying Tensor-based Morphometry to Parametric Surfaces can Improve MRI-based Disease Diagnosis , NeuroImage, 74, 2013, pp. 209-230.
- Wang Y, Chan TF, Toga AW, Thompson PM, “Multivariate Tensor-based Brain Anatomical Surface Morphometry via Holomorphic One-Forms”, 12th International Conference on Medical Image Computing and Computer Assisted Intervention – MICCAI 2009, London, UK, Sep. 2009, pp. 337-344
- Y. Wang, X. Gu, T.F. Chan, P.M. Thompson and S.-T. Yau, “Conformal Slit Mapping and Its Applications to Brain Surface Parameterization“, International Conference on Medical Image Computing and Computer Assisted Intervention – MICCAI 2008, LNCS 5241, pp. 585-593