Studying APOE-e4 Allele Dose Effects with a Univariate Morphometry Biomarker
Gang Wang, Wenju Zhou, Deping Kong, Zongshuai Qu, Maowen Ba, Jinguang Hao, Tao Yao, Qunxi Dong, Yi Su, Eric M Reiman, Richard J Caselli, Kewei Chen, Yalin Wang
Abstract
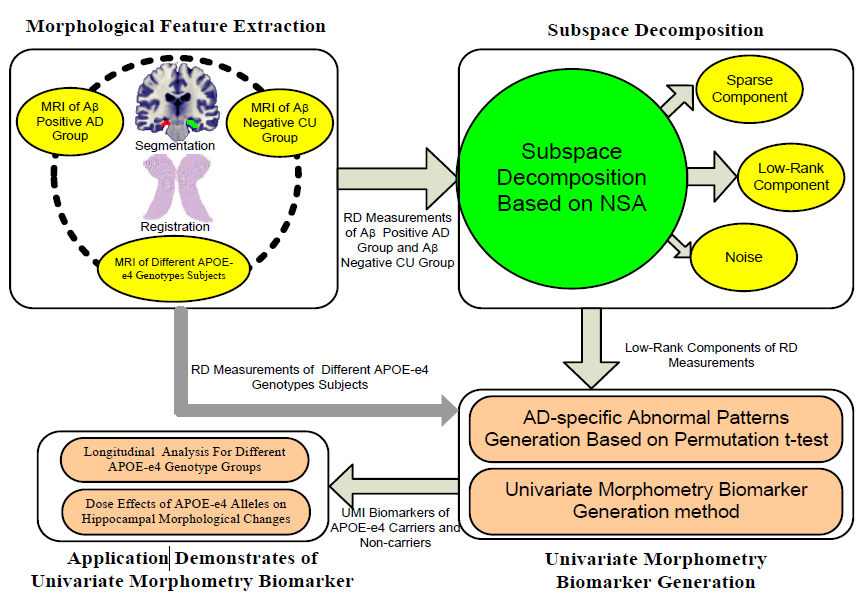
This paper proposes a subspace decomposition method capable of exploiting a univariate morphometry biomarker to represent the hippocampal morphological changes associated with APOE-e4 genetic risk for Alzheimer’s disease (AD) based on magnetic resonance imaging (MRI) data of the elderly cognitively unimpaired (CU) individuals. Rank minimization and sparsity constraint theory drive our method. Rank minimization mechanism combined with improved partial singular value decomposition (SVD) is used to extract group common structures and reduce the computational cost of SVD of the large-scale matrix. Further, sparse constraint considering the local continuity of the hippocampal atrophy regions is applied to recover individual morphological differences. Based on the group common structures of amyloid-β (Aβ) positive AD patients and Aβ negative CU subjects, we identified the regions-of-interest (ROI), which reflect significant morphometry changes caused by the AD development. Then univariate morphometry index (UMI) is constructed from these ROIs. Capitalizing on the study of a large number of cognitively unimpaired late middle-aged and older adults with two, one, and no APOE-e4 alleles, we set up three experiments to study the differences of hippocampal morphological changes among the longitudinal APOE-e4 homozygotes (HM, N=30), heterozygotes (HT, N=49) and non-carriers (NC, N=61). The results show that the proposed UMI enjoys improved computational efficiency comparing to our prior work and demonstrates a more vital ability to identify APOE-e4 dose effects than the raw surface and volume measurements. Our work may provide a cost-effective UMI that helps evaluate AD burden, progression, and response to interventions, potentially facilitating AD clinical trial enrollments.