Signed graph representation learning for functional-to-structural brain network mapping
Tang H, Guo L, Fu X, Wang Y, Mackin S, Ajilore O, Leow AD, Thompson PM, Huang H, Zhan L
Abstract
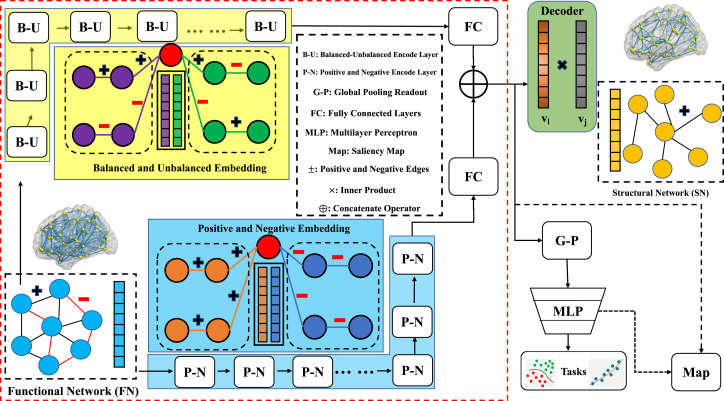
MRI-derived brain networks have been widely used to understand functional and structural interactions among brain regions, and factors that affect them, such as brain development and diseases. Graph mining on brain networks can facilitate the discovery of novel biomarkers for clinical phenotypes and neurodegenerative diseases. Since brain functional and structural networks describe the brain topology from different perspectives, exploring a representation that combines these cross-modality brain networks has significant clinical implications. Most current studies aim to extract a fused representation by projecting the structural network to the functional counterpart. Since the functional network is dynamic and the structural network is static, mapping a static object to a dynamic object may not be optimal. However, mapping in the opposite direction (i.e., from functional to structural networks) are suffered from the challenges introduced by negative links within signed graphs. Here, we propose a novel graph learning framework, named as Deep Signed Brain Graph Mining or DSBGM, with a signed graph encoder that, from an opposite perspective, learns the cross-modality representations by projecting the functional network to the structural counterpart. We validate our framework on clinical phenotype and neurodegenerative disease prediction tasks using two independent, publicly available datasets (HCP and OASIS). Our experimental results clearly demonstrate the advantages of our model compared to several state-of-the-art methods.