Developing Univariate Neurodegeneration Biomarkers with Low-Rank and Sparse Subspace Decomposition
Wang G, Dong Q, Wu J, Su Y, Chen K, Su Q, Zhang X, Hao J, Yao T, Liu L, Zhang C, Caselli RJ, Reiman EM, Wang Y, ADNI Group
Abstract
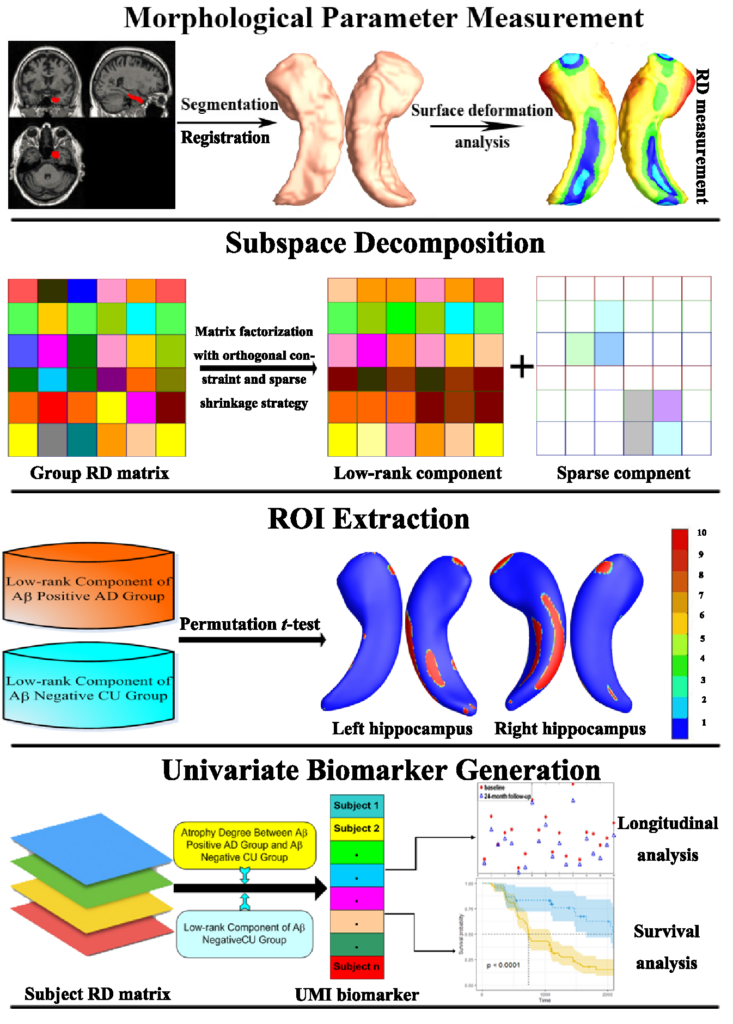
Cognitive decline due to Alzheimer’s disease (AD) is closely associated with brain structure alterations captured by structural magnetic resonance imaging (sMRI). It supports the validity to develop sMRI-based univariate neurodegeneration biomarkers (UNB). However, existing UNB work either fails to model large group variances or does not capture AD dementia (ADD) induced changes. We propose a novel low-rank and sparse subspace decomposition method capable of stably quantifying the morphological changes induced by ADD. Specifically, we propose a numerically efficient rank minimization mechanism to extract group common structure and impose regularization constraints to encode the original 3D morphometry connectivity. Further, we generate regions-of-interest (ROI) with group difference study between common subspaces of Aβ+ AD and Aβ- cognitively unimpaired (CU) groups. A univariate morphometry index (UMI) is constructed from these ROIs by summarizing individual morphological characteristics weighted by normalized difference between Aβ+ AD and Aβ- CU groups. We use hippocampal surface radial distance feature to compute the UMIs and validate our work in the Alzheimer’s Disease Neuroimaging Initiative (ADNI) cohort. Our experimental results outperform traditional hippocampal volume measures and suggest the application of UMI as a potential UNB.