Deep Multimodal Brain Network Learning for Joint Analysis of Structural Morphometry and Functional Connectivity
Wen Zhang, Yalin Wang
Abstract
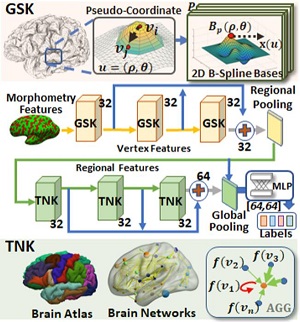
Learning from the multimodal brain imaging data attracts a large amount of attention in medical image analysis due to the proliferation of multimodal data collection. It is widely accepted that multimodal data can provide complementary information than mining from a single modality. However, unifying the image-based knowledge from the multimodal data is very challenging due to different image signals, resolution, data structure, etc.. In this study, we design a supervised deep model to jointly analyze brain morphometry and functional connectivity on the cortical surface and we name it deep multimodal brain network learning (DMBNL). Two graph-based kernels, i.e., geometry-aware surface kernel (GSK) and topology-aware network kernel (TNK), are proposed for processing the cortical surface morphometry and brain functional network. The vertex features on the cortical surface from GSK is pooled and feed into TNK as its initial regional features. In the end, the graph-level feature is computed for each individual and thus can be applied for classification tasks. We test our model on a large autism imaging dataset. The experimental results prove the effectiveness of our model.
Figures (click on each for a larger version):
Related Publications
- Zhang W, Zhan L, Thompson PM, Wang Y,, “Deep Representation Learning For Multimodal Brain Networks”, 23rd International Conference on Medical Image Computing and Computer Assisted Intervention – MICCAI, Lima, Peru, Oct. 2020